Overview¶
A high-performance topological machine learning toolbox in Python
giotto-tda is a high performance topological machine learning toolbox in Python built on top of
scikit-learn and is distributed under the GNU AGPLv3 license. It is part of the Giotto family of open-source projects.
Guiding principles¶
scikit-learnStrictly adhere to thescikit-learnAPI and development guidelines, inherit the strengths of that framework.joblib).
30s guide to giotto-tda¶
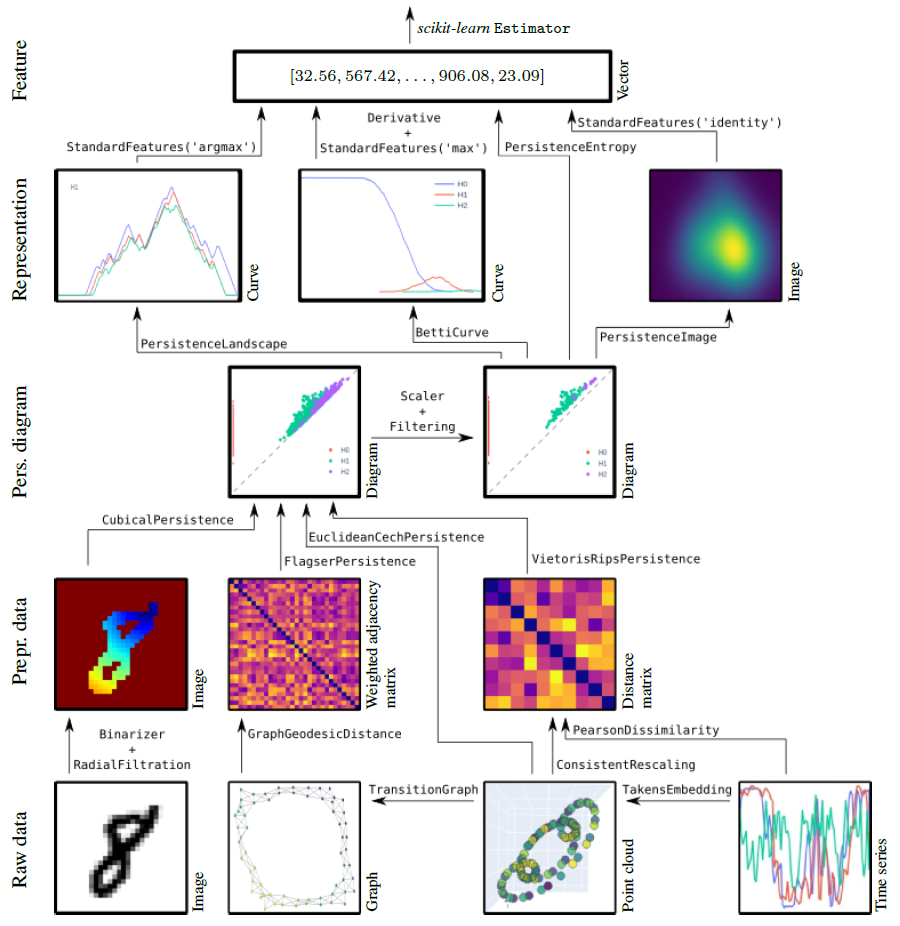
For installation instructions, see the installation instructions.
The functionalities of giotto-tda are provided in scikit-learn–style transformers.
This allows you to generate topological features from your data in a familiar way. Here is an example with the VietorisRipsPersistence transformer:
from gtda.homology import VietorisRipsPersistence
VR = VietorisRipsPersistence()
which computes topological summaries, called persistence diagrams, from collections of point clouds or weighted graphs, as follows:
diagrams = VR.fit_transform(point_clouds)
A plotting API allows for quick visual inspection of the outputs of many of giotto-tda’s transformers. To visualize the i-th output sample, run
diagrams = VR.plot(diagrams, sample=i)
You can create scalar or vector features from persistence diagrams using giotto-tda’s dedicated transformers. Here is an example with the PersistenceEntropy transformer:
from gtda.diagrams import PersistenceEntropy
PE = PersistenceEntropy()
features = PE.fit_transform(diagrams)
features is a two-dimensional numpy array. This is important to making this type of topological feature generation fit into a typical machine learning workflow from scikit-learn.
In particular, topological feature creation steps can be fed to or used alongside models from scikit-learn, creating end-to-end pipelines which can be evaluated in cross-validation,
optimised via grid-searches, etc.:
from sklearn.ensemble import RandomForestClassifier
from gtda.pipeline import make_pipeline
from sklearn.model_selection import train_test_split
X_train, X_valid, y_train, y_valid = train_test_split(point_clouds, labels)
RFC = RandomForestClassifier()
model = make_pipeline(VR, PE, RFC)
model.fit(X_train, y_train)
model.score(X_valid, y_valid)
giotto-tda also implements the Mapper algorithm as a highly customisable scikit-learn Pipeline, and provides simple plotting functions for visualizing output Mapper graphs and have real-time interaction with the pipeline parameters:
from gtda.mapper import make_mapper_pipeline
from sklearn.decomposition import PCA
from sklearn.cluster import DBSCAN
pipe = make_mapper_pipeline(filter_func=PCA(), clusterer=DBSCAN())
plot_interactive_mapper_graph(pipe, data)
Resources¶
Tutorials and examples¶
We provide a number of tutorials and examples, which offer:
quick start guides to the API;
in-depth examples showcasing more of the library’s features;
intuitive explanations of topological techniques.
Use cases¶
A selection of use cases for giotto-tda is collected at this page.
Please note, however, that some of these were written for past versions of giotto-tda. In some cases,
only small modifications are needed to run them on recent versions, while in others it is best to install
the relevant past version of giotto-tda (preferably in a fresh environmnent). In a couple of cases,
the legacy giotto-learn or giotto-learn-nightly will be needed.
What’s new¶
Major Features and Improvements¶
An object-oriented API for interactive plotting of Mapper graphs has been added with the
MapperInteractivePlotter(#586). This is intended to supersedeplot_interactive_mappergraph as it allows for inspection of the current state of the objects change by interactivity. See also “Backwards-Incompatible Changes” below.Further citations have been added to the mathematical glossary (#564).
Bug Fixes¶
A bug preventing
EuclideanCechPersistencefrom working correctly on point clouds in more than 2 dimensions has been fixed (#588).A validation bug preventing
VietorisRipsPersistenceandWeightedRipsPersistencefrom accepting non-empty dictionaries asmetric_paramshas been fixed (#590).A bug causing an exception to be raised when
node_color_statisticwas passed as a numpy array inplot_static_mapper_graphhas been fixed (#576).
Backwards-Incompatible Changes¶
A major change to the behaviour of the (static and interactive) Mapper plotting functions
plot_static_mapper_graphandplot_interactive_mapper_graphwas introduced in #584. The newMapperInteractivePlotterclass (see “Major Features and Improvements” above) also follows this new API. The main changes are as follows:color_by_columns_dropdownhas been eliminated.color_variablehas been renamed tocolor_features(but cannot be an array).An additional keyword argument
color_datahas been added to more clearly separate the inputdatato the Mapper pipeline from the data to be used for coloring.node_color_statisticis now applied column by column – previously it could end up being applied to 2d arrays as a whole.The defaults for color-related arguments lead to index values instead of the mean of the data.
The default for
weight_paramsinWeightedRipsPersistenceis now the empty dictionary, andNoneis no longer allowed (#595).
Thanks to our Contributors¶
This release contains contributions from many people:
Umberto Lupo, Wojciech Reise, Julian Burella Pérez, Sean Law, Anibal Medina-Mardones, and Lewis Tunstall
We are also grateful to all who filed issues or helped resolve them, asked and answered questions, and were part of inspiring discussions.